Abstract
A polka-dotted fruit fly, Drosophila guttifera, has a unique pigmentation pattern made of black melanin and serves as a good model system to study color pattern formation. Because of its short generation time and the availability of transgenics, it is suitable for dissecting the genetic mechanisms of color pattern formation. While the ecology and life history of D. guttifera in the wild are not well understood, it is known to be resistant to a mushroom toxin, and this physiological trait is under molecular scrutiny. Pigmentation around crossveins and longitudinal vein tips is common in closely related species of the quinaria group, in addition to which D. guttifera has evolved species-specific pigmentation spots around the campaniform sensilla. Regulatory evolution of the Wnt signaling ligand Wingless, which locally induces pigmentation in the developing wing epithelium, has driven the evolution of distinct aspects of wing and body pigmentation. A melanin biosynthesis pathway gene, yellow, is also involved in the elaboration of these traits, downstream of wingless. Unraveling the detailed mechanism of pigmentation pattern formation of this species sheds light on the general principles of morphological evolution and foreshadows potential parallels with other systems, such as the pigmented wings of butterflies.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Drosophila guttifera
- Pigmentation
- Color pattern
- Evolution
- Development
- Transgenic
- Cis-regulatory element
- Phylogeny
- Ecology
- Life history
- Taxonomy
1 Introduction
Research on butterfly color patterns has greatly advanced in recent years. Knowledge of the characteristics of the genome, mechanisms of pattern formation, and the function and evolutionary mode of the pattern is rapidly growing. This was enabled by utilization of multiple model species, including species of Bicyclus, Heliconius, Junonia, Vanessa, Papilio, and others, and by the best use of characteristics of materials (Nijhout 1991; Carroll et al. 1994; Brakefield et al. 1996; Joron et al. 2011; Reed et al. 2011; The Heliconius Genome Consortium 2012; Martin et al. 2012; Kunte et al. 2014; Monteiro 2015; Nishikawa et al. 2015; Beldade and Peralta 2017).
In vertebrates, zebrafish (Danio rerio) has been a model of color pattern formation, and recently, domestic and wild cats and a four-striped mouse (Rhabdomys pumilio) were also used for research, making this an exciting time for color pattern studies (Singh and Nüsslein-Volhard 2015; Kaelin et al. 2012; Mallarino et al. 2016).
We have been using a dipteran insect, Drosophila guttifera, to study a mechanism of color pattern formation (Fig. 16.1). D. guttifera has a pattern on its wings, which is a commonality with butterflies; however, there are also some important differences. In contrast with the pigmented scales of butterflies and moths (as an exception, see Stavenga et al. 2010), Drosophila pigmentation is embedded in the cuticle layers of the wing membrane. This pigmentation is believed to be made of black melanin. A congeneric species, Drosophila melanogaster, is a model organism widely used in genetics and various biological researches, and we can utilize its knowledge, techniques, and resources to study D. guttifera. This phylogenetic proximity is an asset, as it is possible to transfer a part of the genetic system, such as an enhancer involved in pattern formation, into D. melanogaster and analyze its function in a heterologous context. D. guttifera has the potential to approach the same problem of color pattern as in butterflies but from a different angle. It also enables a good comparison, since its complex pigmentation patterns evolved independently from the ones seen in butterflies.
In this chapter, we present an overview of the biology of D. guttifera. Then we discuss differences in pattern formation between D. guttifera and butterflies and the advantage and potential of D. guttifera to contribute to the general understanding of animal color pattern formation.
2 Phylogenetic Position of D. guttifera
Fruit flies (drosophilid flies) belong to family Drosophilidae, order Diptera, and consist of 72 genera and more than 4000 described species (Yassin 2013). Among them, genus Drosophila includes more than 1160 described species (Markow and O’Grady 2006; Toda 2017). The best-studied species, D. melanogaster, also belongs to this genus. It should be noted, however, that the genus Drosophila is not monophyletic and potentially includes multiple genera within this clade, and there is ongoing debate on the proper taxonomic treatment of this genus (O’Grady 2010).
D. guttifera was described by an English entomologist, Francis Walker, based on a specimen collected in Florida (Walker 1849). This description consisted of 4 lines in Latin and 21 lines in English with no illustration and was one of many descriptions of a museum collection of the British Museum. In his taxonomic revision of North American drosophilids, Sturtevant (1921) examined multiple specimens of D. guttifera and redescribed the morphological features. Sturtevant (1942) established “species groups” to classify species within the genus Drosophila. D. guttifera was assigned to a monospecific guttifera group. He also established the quinaria group, which includes 11 species (D. quinaria, deflecta, palustris, subpalustris, occidentalis, suboccidentalis, munda, subquinaria, transversa, and possibly phalerata and nigromaculata). Patterson (1943) revised drosophilids of the Southwestern United States and Northern Mexico and redescribed many species with beautiful illustrations. D. guttifera was redescribed with illustrations of a pupa and internal organs of reproduction and a color illustration of the whole body. Patterson also described three new species in the quinaria group (D. suffusca, tenebrosa, and innubila). After that, many species were described in the quinaria group, and currently it includes 31 species (Markow and O’Grady 2006, Toda 2017).
The close relationship between D. guttifera and the quinaria group is almost certain at this time, based on molecular genetic evidence (Perlman et al. 2003; Izumitani et al. 2016). Morphological similarity between D. guttifera and the quinaria group was also noticed (Patterson and Stone 1952), and some authors even placed D. guttifera in the quinaria group (Throckmorton 1962, 1975; Markow and O’Grady 2006). Species-level relationships among D. guttifera and species of the quinaria group are not completely resolved; however, the commonly supported result is bifurcation into two clades, one including mostly North American species and one including mostly Eurasian species (Perlman et al. 2003, Markow and O’Grady 2006, Izumitani et al. 2016).
There are species with pigment patterns on the thorax, abdomen, and wings to various degrees in the quinaria group (Patterson 1943; Werner and Jaenike 2017), but D. guttifera has distinctive vertical stripes on the thorax and a polka dot pattern on the abdomen and wings. Even when compared with the quinaria group species, D. guttifera has the most prominently pigmented appearance.
3 Food Habits, Poison Resistance, and Behavioral Ecology of D. guttifera
The life history and ecology of D. guttifera in the wild have not been well studied. There are many species of the quinaria group that utilize mushrooms as a food source. Sturtevant (1921) assumed D. guttifera is also a mushroom feeder based on the facts that D. guttifera was found around mushrooms and that he could rear D. guttifera, from eggs to adults, with mushrooms (he noted that both gill fungi and pore fungi can be utilized, but he did not describe mushroom species). Bunyard and Foote (1990a) studied what kind of dipteran insects emerged from mushrooms collected in the state of Ohio and reported that D. guttifera emerged from two mushroom species, Psilocybe polytrichophila and Collybia dryophila. They tested oviposition site preference among commercial Agaricus bisporus, banana, tomato, lettuce, and agar and found that Agaricus was the most preferred site (Bunyard and Foote 1990b). They also confirmed that D. guttifera can grow from eggs to adults with Agaricus. In laboratory conditions, however, we can keep strains of D. guttifera with artificial food containing sugar/corn meal/yeast/agar (sugar food) or molasses/corn meal/yeast/agar (molasses food) without adding mushrooms.
Some fungus-feeding drosophilids are known to have high tolerance to a mushroom toxin, alpha-amanitin, which is highly poisonous to most animals (Spicer and Jaenike 1996). D. guttifera has the potential to be a model system to study this phenomenon. Alpha-amanitin exerts its toxicity by binding to RNA polymerase II, an enzyme essential for transcription. A mutant strain of D. melanogaster with high alpha-amanitin tolerance had an amino acid substitution in RNA polymerase II (Chen et al. 1993). However, D. guttifera and other species with the tolerance do not have the same substitution, indicating that other mechanisms are involved (Stump et al. 2011). There are other strains of D. melanogaster with alpha-amanitin tolerance but without RNA polymerase II mutation. The responsible locus was mapped, and gene expression profiles were analyzed in these strains (Begum and Whitley 2000; Mitchell et al. 2014, 2015).
There are some other studies of D. guttifera behavior. Oviposition site preference of D. guttifera was affected by larval food condition, and this is known as a classic example of olfactory conditioning of animals (Cushing 1941). The mating behavior of D. guttifera was also studied (Grossfield 1977). The ecological significance and function of pigmentation patterns of D. guttifera is not well understood. Some drosophilids are known to use wing pigmentation in courtship displays (Ringo and Hodosh 1978; Yeh et al. 2006; Fuyama 1979). Dombeck and Jaenike (2004) analyzed fitness effects of abdominal spot number in D. falleni.
4 The Evolution of Wing Pigmentation Pattern
Dombeck and Jaenike (2004) analyzed and illustrated the evolutionary path of wing and abdominal pigmentations of D. guttifera and seven species of the quinaria group. We summarize here the evolution of wing pigmentation pattern of D. guttifera and the quinaria group species based on molecular phylogenetics (Fig. 16.2). As previously explained, the quinaria group is divided into two major clades (Perlman et al. 2003; Markow and O’Grady 2006, Izumitani et al. 2016). We defined the clade with mostly North American species as “clade A” and the clade with mostly Eurasian species as “clade B.” Species in clade A have relatively simple patterns; pigmentations are formed only around crossveins except in D. innubila, which has no pigmentation. The evolution of patterns in clade B is rather complicated. The relationships among basal species of clade B [D. guttifera, nigromaculata, and (deflecta + palustris + subpalustris)] have not been completely resolved, because the topologies of the phylogenetic trees depend on the analytical methods. These four species have pigmentations around crossveins and longitudinal vein tips. In addition, D. guttifera has pigmentations around the campaniform sensilla, which is unique to this species [at least unique among the clade of (quinaria group + D. guttifera) and probably among the genus Drosophila]. Among the rest of the species in clade B, D. quinaria has weak pigmentations on the tips of longitudinal veins in addition to crossveins. D. recens and many other species within this cluster have pigmentations around crossveins. D. kuntzei, which has a similar pattern to D. quinaria, branches from the most basal position of clade B according to Perlman et al. (2003), although the statistical support for this topology was low. Due to the lack of a robust phylogeny, it would be premature to propose a simple scenario stepwise pattern of gain and loss within the quinaria group. It is plausible that the instances of longitudinal vein tip pigmentation are the result of convergent evolution, perhaps via parallel mechanisms, although we cannot exclude the possibility of a single gain of the longitudinal vein tip pigmentation and a secondary loss in derived species of clade B. Nevertheless, the other dot-like patterns of D. guttifera, which overlap in position with innervated cupules known as campaniform sensilla (see below), are unique to this species and are assumed to form a true evolutionary novelty.
5 Wing Pigmentation Pattern Formation in Drosophila
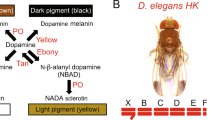
The initial study of the mechanism of wing pigmentation pattern formation was done by True et al. (1999). They argued that patterns are formed through patterning by gene expression and subsequent elaboration by precursor trafficking through wing veins, based on experiments using Drosophila grimshawi (synonym of Idiomyia grimshawi), D. rajasekari (synonym of D. biarmipes), and mutants and transgenics of D. melanogaster. Wittkopp et al. (2002) studied the function of yellow and ebony genes in the body trunk and wings of D. melanogaster. They also showed that the future spot position had more Yellow protein and less Ebony protein. Yellow is known to enhance black melanin synthesis, and Ebony is an enzyme that conjugates beta-alanine to dopamine and produces NBAD (N-beta-alanyldopamine) resulting in repression of black melanin synthesis. Gompel et al. (2005) analyzed the regulation of yellow gene expression in D. biarmipes and showed that evolution of an enhancer (a sequence that enhances expression of a nearby gene) was involved in the gain of pigmentation. In D. biarmipes and D. guttifera, they showed that Yellow protein was localized in future black spots and Ebony protein was localized in future transparent (no pigmentation) places. The yellow expression in the anteriodistal part of the wing in D. biarmipes results from regulation by at least two factors: posterior expression of engrailed repressing the yellow expression and anteriodistal expression of Distal-less enhancing expressions of yellow and other pigmentation genes (Gompel et al. 2005; Arnoult et al. 2013).
6 Features of Wing Pigmentation Pattern in D. guttifera
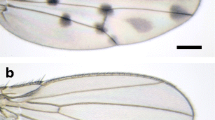
D. guttifera has prominent black polka dots on its wings, and these are believed to be made with melanin (Fig. 16.3). Pigmentations are formed around crossveins, longitudinal vein tips, and the campaniform sensilla. Weak pigmentations are also formed in intervein regions. As mentioned previously, crossvein pigmentation is widely observed in the quinaria group and also found in many species in other species groups. The crossvein pigmentation in D. guttifera is constricted in the center, forming an hourglass shape (or calabash shape), and this is unique to this species. Longitudinal vein tip pigmentations are observed in a few species, but the pigmentation area is largest in D. guttifera. Campaniform sensilla pigmentation is a trait unique to D. guttifera, although some species, such as a Hawaiian species, Idiomyia grimshawi (synonym of Drosophila grimshawi), have dappled spots all over the wings. The campaniform sensilla are lined on the third longitudinal vein in the same way as in other drosophilids, but in D. guttifera, one campaniform sensillum is also found on the fifth longitudinal vein, which is unique to this species. This campaniform sensillum is also surrounded by pigmentation (Sturtevant 1921; Werner et al. 2010). The wing pigmentation of D. guttifera starts to form in the pupal period, and it continues until one day old adult (Fukutomi et al. 2017).
7 Wingless Gene Induces Pigmentation Pattern Formation in D. guttifera
Werner et al. (2010) analyzed the cis-regulatory region of the yellow gene and identified vein spot CRE, which is an enhancer driving expression in all the polka dots, and intervein shade CRE, which is an enhancer driving expression in the intervein region. Vein spot CRE drove polka dots in D. guttifera but drove around crossveins and longitudinal vein tips if introduced in D. melanogaster. This difference means there is a difference in localization of a trans-regulatory factor that has an input to vein spot CRE. Gene expression patterns were known for several genes in D. melanogaster, and therefore they found candidate genes from genes showing similar expression with the vein spot CRE pattern. Among the candidate genes, wingless, a gene encoding a ligand of the Wnt signaling pathway, showed expression in the center of future spot positions (crossveins, longitudinal vein tips, and campaniform sensilla) in D. guttifera. There was no wingless expression in the campaniform sensilla in a closely related species, D. deflecta, which does not have pigmentation around them. A spontaneous mutant line of D. guttifera, schwarzvier, has additional pigmentation on the fourth longitudinal vein. In this mutant line, wingless was ectopically expressed on the fourth longitudinal vein. To obtain direct functional evidence, they tried to make ectopic expressions of wingless by construction of the GAL4/UAS system in D. guttifera. Although they did not obtain optimal GAL4 lines, they found that one of the UAS-wingless lines had ectopic expression of wingless, probably caused by the enhancer trap mechanism. In this line, wingless was expressed ectopically on the second, third, and fourth longitudinal veins of pupal wings, and additional pigmentation was formed on these veins in adult wings. With these evidences, they concluded that wingless is the upstream trans-factor that induces pigmentation.
In Heliconius and Limenitis butterflies, the WntA gene, which also seems to encode a ligand of Wnt signaling, is involved in specifying wing pattern shapes, including in melanic elements (Martin et al. 2012; Gallant et al. 2014; Martin and Reed 2014). In Junonia coenia and some other butterfly species, wingless is known to be expressed in future pattern elements called basal (B), discal (DI and DII), and marginal (EI) elements (Carroll et al. 1994; Martin and Reed 2010, 2014; Huber et al. 2015) and was also identified at the center of eyespot patterns (Monteiro et al. 2006). The thoracic pattern of larval Bombyx mori is also regulated by Wnt1 (homolog of wingless) (Yamaguchi et al. 2013). Evolutionary roles of secreted ligand genes such as wingless are reviewed in chapter 4 of this book (Martin and Courtier-Orgogozo 2017).
Werner et al. (2010) proposed a model of pigmentation pattern formation based on the assumption that Wingless protein diffuses from the source and serves as a long-range signal. There are a limited number of cells expressing wingless, and they are located in centers of future pigmented spots. In their model, secreted Wingless protein is diffused or transported to wider regions and transduces the signal. The signal is probably mediated by an unknown transcription factor and activates transcription of melanin synthesis-related genes, including yellow. Melanin should be synthesized by products of these genes and wings are consequently pigmented. This model should be validated by future research.
8 Cis-Regulatory Evolution of Wingless
The expression pattern of wingless evolved uniquely in D. guttifera. To examine how this unique expression pattern evolved, the genomic region around wingless was analyzed using a fluorescent reporter assay. As a result, three novel enhancer activities (in longitudinal vein tips, campaniform sensilla, and thoracic stripes) were found (Fig. 16.4). These novel enhancer activities are thought to have been involved in the evolution of the novel pigmentation pattern (Koshikawa et al. 2015). This study provided unique insights into the evolution of novel traits, illustrating how gains of novel enhancer activities at developmental regulatory gene were associated with derived expression domains and the emergence of novel traits (Rebeiz et al. 2011; Koshikawa et al. 2015; Rebeiz and Williams 2017).
Enhancers driving pupal wing and thoracic expressions of wingless in D. guttifera. Color code indicates correspondence of enhancer positions and expression domains. Green: wing margin. Blue: crossveins. Purple: longitudinal vein tips. Yellow: campaniform sensilla. Brown: thoracic stripes. Expressions in the wing margin and crossveins are ancestral (common in D. melanogaster and D. guttifera), and the longitudinal vein tips, campaniform sensilla, and thoracic stripes are novel (found in D. guttifera but not in D. melanogaster) (Modified from Koshikawa et al. (2015) and Koshikawa (2015))
We can generalize this concept as follows. In many organisms, gains of novel expression domains by gains of enhancer activities for a developmental regulatory gene could be a part of possible mechanisms of heterotopy (evolutionary duplication of a pre-existing trait in a different place on the body) (Gould 1977; West-Eberhard 2003; Rubinstein and de Souza 2013; Rebeiz et al. 2015; for more discussion see Koshikawa 2015).
9 Trials of Artificial Production of Pigmentation on D. melanogaster Wings
For now, only two genes, the upstream pattern inducer wingless and the melanin synthesis-related gene yellow, have been identified in the machinery required for pigmentation pattern formation in D. guttifera. In many cases, the Wingless signal is transduced through the so-called canonical pathway, where Pangolin/dTCF is an effector transcription factor regulating transcriptions of downstream genes. There were consensus sequences of Pangolin/dTCF binding sites in vein spot CRE, but replacement of these sequences by nonsense sequences did not change the expression pattern of the reporter gene (Werner et al. 2010). This means that the positional information of wingless does not directly regulate yellow through the canonical Wnt pathway. Involvement of another transcription factor is assumed, but so far it has not been identified. Furthermore, we know yellow is involved in pigmentation, but overexpression of yellow alone does not cause additional pigmentation in D. melanogaster (Gompel et al. 2005; Riedel et al. 2011). Proper expression or repression of melanin synthesis-related genes and/or proper supply of melanin precursors, such as dopa and dopamine, could be required for artificial production of pigmentation in D. melanogaster wings.
10 Diversity and Generality in Color Pattern Formation
We summarized above what was revealed by studies of D. guttifera, but will it apply to pattern formation in other organisms? Due to the experimental strengths of this system, we can be optimistic that we will reach an integrated model for pigmentation pattern formation in Drosophila. Butterflies show interesting parallels with the Drosophila wing patterning genes, as Wnt genes and Distal-less are key players in both lineages (Werner et al. 2010; Martin et al. 2012; Brakefield et al. 1996; Arnoult et al. 2013). If we expand the comparison to vertebrates, there are large differences in genes involved in pattern formation and melanin synthesis (Kopp 2009; Kronforst et al. 2012; Kaelin et al. 2012; Mallarino et al. 2016). Still we assume we can find some common mechanisms, such as a way of measuring distance in a tissue, and a hierarchical regulatory architecture. Comparing comprehensive datasets will be instrumental in answering this question of fundamental interest for our understanding of the mechanisms that generate biodiversity on Earth.
References
Arnoult L, Su KF, Manoel D, Minervino C, Magriña J, Gompel N, Prud’homme B (2013) Emergence and diversification of fly pigmentation through evolution of a gene regulatory module. Science 339(6126):1423–1426. doi:10.1126/science.1233749
Begun DJ, Whitley P (2000) Genetics of alpha-amanitin resistance in a natural population of Drosophila melanogaster. Heredity 85(2):184–190. doi:10.1046/j.1365-2540.2000.00729.x
Beldade P, Peralta CM (2017) Developmental and evolutionary mechanisms shaping butterfly eyespots. Curr Opin Insect Sci 19:22–29. doi:10.1016/j.cois.2016.10.006
Brakefield PM, Gates J, Keys D, Kesbeke F, Wijngaarden PJ, Monteiro A, French V, Carroll SB (1996) Development, plasticity and evolution of butterfly eyespot patterns. Nature 384(6606):236–242. doi:10.1038/384236a0
Bunyard B, Foote BA (1990a) Acalyptrate Diptera reared from higher fungi in northeastern Ohio. Entomol News 101(2):117–121
Bunyard B, Foote BA (1990b) Biological notes on Drosophila guttifera (Diptera: Drosophilidae), a consumer of mushrooms. Entomol News 101(3):161–163
Carroll SB, Gates J, Keys DN, Paddock SW, Panganiban GE, Selegue JE, Williams JA (1994) Pattern formation and eyespot determination in butterfly wings. Science 265(5168):109–114. doi:10.1126/science.7912449
Chen Y, Weeks J, Mortin MA, Greenleaf AL (1993) Mapping mutations in genes encoding the two large subunits of Drosophila RNA polymerase II defines domains essential for basic transcription functions and for proper expression of developmental genes. Mol Cell Biol 13(7):4214–4222. doi:10.1128/MCB.13.7.4214
Cushing JE (1941) An experiment in olfactory conditioning in Drosophila guttifera. Proc Natl Aca Sci U S A 27(11):496–499
Dombeck I, Jaenike J (2004) Ecological genetics of abdominal pigmentation in Drosophila falleni. Evolution 58(3):587–596. doi:10.1554/03-299
Fukutomi Y, Matsumoto K, Agata K, Funayama N, Koshikawa S (2017) Pupal development and pigmentation process of a polka-dotted fruit fly, Drosophila guttifera (Insecta, Diptera). Dev Genes Evol 227(3):171–180. doi:10.1007/s00427-017-0578-3
Fuyama Y (1979) A visual stimulus in the courtship of Drosophila suzukii. Experientia 35(10):1327–1328
Gallant JR, Imhoff VE, Martin A, Savage WK, Chamberlain NL, Pote BL, Peterson C, Smith GE, Evans B, Reed RD, Kronforst MR, Mullen SP (2014) Ancient homology underlies adaptive mimetic diversity across butterflies. Nat Commun 5:4817. doi:10.1038/ncomms5817
Gompel N, Prud’homme B, Wittkopp PJ, Kassner VA, Carroll SB (2005) Chance caught on the wing: cis-regulatory evolution and the origin of pigment patterns in Drosophila. Nature 433(7025):481–487. doi:10.1038/nature03235
Gould SJ (1977) Ontogeny and phylogeny. Harvard University Press, Cambridge
Grossfield J (1977) Drosophila courtship: decapitated quinaria group females. J NY Entomol Soc 85(3):119–126
Huber B, Whibley A, Poul YL, Navarro N, Martin A, Baxter S, Shah A, Gilles B, Wirth T, McMillan WO, Joron M (2015) Conservatism and novelty in the genetic architecture of adaptation in Heliconius butterflies. Heredity 114(5):515–524. doi:10.1038/hdy.2015.22
Izumitani HF, Kusaka Y, Koshikawa S, Toda MJ, Katoh T (2016) Phylogeography of the subgenus Drosophila (Diptera: Drosophilidae): evolutionary history of faunal divergence between the old and the new worlds. PLoS One 11(7), e0160051. doi:10.1371/journal.pone.0160051
Joron M, Frezal L, Jones RT, Chamberlain NL, Lee SF, Haag CR, Whibley A, Becuwe M, Baxter SW, Ferguson L, Wilkinson PA, Salazar C, Davidson C, Clark R, Quail MA, Beasley H, Glithero R, Lloyd C, Sims S, Jones MC, Rogers J, Jiggins CD, ffrench-Constant RH (2011) Chromosomal rearrangements maintain a polymorphic supergene controlling butterfly mimicry. Nature 477(7363):203–206. doi:10.1038/nature10341
Kaelin CB, Xu X, Hong LZ, David VA, McGowan KA, Schmidt-Küntzel A, Roelke ME, Pino J, Pontius J, Cooper GM, Manuel H, Swanson WF, Marker L, Harper CK, van Dyk A, Yue B, Mullikin JC, Warren WC, Eizirik E, Kos L, O’Brien SJ, Barsh GS, Menotti-Raymond M (2012) Specifying and sustaining pigmentation patterns in domestic and wild cats. Science 337(6101):1536–1541. doi:10.1126/science.1220893
Kopp A (2009) Metamodels and phylogenetic replication: a systematic approach to the evolution of developmental pathways. Evolution 63(11):2771–2789. doi:10.1111/j.1558-5646.2009.00761.x
Koshikawa S (2015) Enhancer modularity and the evolution of new traits. Fly (Austin) 9(4):155–159. doi:10.1080/19336934.2016.1151129
Koshikawa S, Giorgianni MW, Vaccaro K, Kassner VA, Yoder JH, Werner T, Carroll SB (2015) Gain of cis-regulatory activities underlies novel domains of wingless gene expression in Drosophila. Proc Natl Acad Sci U S A 112(24):7524–7529. doi:10.1073/pnas.1509022112
Kronforst MR, Barsh GS, Kopp A, Mallet J, Monteiro A, Mullen SP, Protas M, Rosenblum EB, Schneider CJ, Hoekstra HE (2012) Unraveling the thread of nature’s tapestry: the genetics of diversity and convergence in animal pigmentation. Pigment Cell Melanoma Res 25(4):411–433. doi:10.1111/j.1755-148X.2012.01014.x
Kunte K, Zhang W, Tenger-Trolander A, Palmer DH, Martin A, Reed RD, Mullen SP, Kronforst MR (2014) Doublesex is a mimicry supergene. Nature 507(7491):229–232. doi:10.1038/nature13112
Mallarino R, Henegar C, Mirasierra M, Manceau M, Schradin C, Vallejo M, Beronja S, Barsh GS, Hoekstra HE (2016) Developmental mechanisms of stripe patterns in rodents. Nature 539(7630):518–523. doi:10.1038/nature20109
Martin A, Reed RD (2010) Wingless and aristaless2 define a developmental ground plan for moth and butterfly wing pattern evolution. Mol Biol Evol 27(12):2864–2878. doi:10.1093/molbev/msq173
Martin A, Papa R, Nadeau NJ, Hill RI, Counterman BA, Halder G, Jiggins CD, Kronforst MR, Long AD, McMillan WO, Reed RD (2012) Diversification of complex butterfly wing patterns by repeated regulatory evolution of a Wnt ligand. Proc Natl Aca Sci U S A 109(31):12632–12637. doi:10.1073/pnas.1204800109
Martin A, Reed RD (2014) Wnt signaling underlies evolution and development of the butterfly wing pattern symmetry systems. Dev Biol 395(2):367–378. doi:10.1016/j.ydbio.2014.08.031
Martin A, Courtier-Orgogozo V (2017) Morphological evolution repeatedly caused by mutations in signaling ligand genes. In: Diversity and evolution of butterfly wing patterns: an integrative approach. Springer, New York
Markow TA, O’Grady PM (2006) Drosophila: a guide to species identification and use. Academic Press, New York
Mitchell CL, Saul MC, Lei L, Wei H, Werner T (2014) The mechanisms underlying α-amanitin resistance in Drosophila melanogaster: a microarray analysis. PLoS One 9(4):e93489. doi:10.1371/journal.pone.0093489
Mitchell CL, Yeager RD, Johnson ZJ, D’Annunzio SE, Vogel KR, Werner T (2015) Long-term resistance of Drosophila melanogaster to the mushroom toxin alpha-amanitin. PLoS One 10(5):e0127569. doi:10.1371/journal.pone.0127569
Monteiro A (2015) Origin, development, and evolution of butterfly eyespots. Annu Rev Entomol 60:253–271. doi:10.1146/annurev-ento-010814-020942
Monteiro A, Glaser G, Stockslager S, Glansdorp N, Ramos D (2006) Comparative insights into questions of lepidopteran wing pattern homology. BMC Dev Biol 6:52. doi:10.1186/1471-213X-6-52
Nijhout HF (1991) The development and evolution of butterfly wing patterns. Smithsonian Institution Press, Washington, DC
Nishikawa H, Iijima T, Kajitani R, Yamaguchi J, Ando T, Suzuki Y, Sugano S, Fujiyama A, Kosugi S, Hirakawa H, Tabata S, Ozaki K, Morimoto H, Ihara K, Obara M, Hori H, Itoh T, Fujiwara H (2015) A genetic mechanism for female-limited Batesian mimicry in Papilio butterfly. Nat Genet 47(4):405–409. doi:10.1038/ng.3241
O’Grady PM (2010) Whither Drosophila? Genetics 185(2):703–705. doi:10.1534/genetics.110.118232
Patterson JT (1943) The Drosophilidae of the Southwest. University of Texas Publication 4313:7–216
Patterson JT, Stone WS (1952) Evolution in the genus Drosophila. Macmillan Company, New York
Perlman SJ, Spicer GS, Shoemaker DD, Jaenike J (2003) Associations between mycophagous Drosophila and their Howardula nematode parasites: a worldwide phylogenetic shuffle. Mol Ecol 12(1):237–249. doi:10.1046/j.1365-294X.2003.01721.x
Rebeiz M, Jikomes N, Kassner VA, Carroll SB (2011) Evolutionary origin of a novel gene expression pattern through co-option of the latent activities of existing regulatory sequences. Proc Natl Acad Sci U S A 108(25):10036–10043. doi:10.1073/pnas.1105937108
Rebeiz M, Patel NH, Hinman VF (2015) Unraveling the tangled skein: the evolution of transcriptional regulatory networks in development. Annu Rev Genomics Hum Genet 16:103–131. doi:10.1146/annurev-genom-091212-153423
Rebeiz M, Williams TM (2017) Using Drosophila pigmentation traits to study the mechanisms of cis-regulatory evolution. Curr Opin Insect Sci 19:1–7. doi:10.1016/j.cois.2016.10.002
Reed RD, Papa R, Martin A, Hines HM, Counterman BA, Pardo-Diaz C, Jiggins CD, Chamberlain NL, Kronforst MR, Chen R, Halder G, Nijhout HF, McMillan WO (2011) optix drives the repeated convergent evolution of butterfly wing pattern mimicry. Science 333(6046):1137–1141. doi:10.1126/science.1208227
Riedel F, Vorkel D, Eaton S (2011) Megalin-dependent yellow endocytosis restricts melanization in the Drosophila cuticle. Development 138(1):149–158. doi:10.1242/dev.056309
Ringo JM, Hodosh RJ (1978) A multivariate analysis of behavioral divergence among closely related species of endemic Hawaiian Drosophila. Evolution 32(2):389–397. doi:10.1111/j.1558-5646.1978.tb00654.x
Rubinstein M, de Souza FSJ (2013) Evolution of transcriptional enhancers and animal diversity. Philos Trans R Soc B 368(1632):20130017–20130017. doi:10.1098/rstb.2013.0017
Singh AP, Nüsslein-Volhard C (2015) Zebrafish stripes as a model for vertebrate colour pattern formation. Curr Biol 25(2):R81–R92. doi:10.1016/j.cub.2014.11.013
Spicer GS, Jaenike J (1996) Phylogenetic analysis of breeding site use and α-amanitin tolerance within the Drosophila quinaria species group. Evolution 50(6):2328–2337. doi:10.2307/2410701
Stavenga DG, Giraldo MA, Leertouwer HL (2010) Butterfly wing colors: glass scales of Graphium sarpedon cause polarized iridescence and enhance blue/green pigment coloration of the wing membrane. J Exp Biol 213(10):1731–1739. doi:10.1242/jeb.041434
Stump AD, Jablonski SE, Bouton L, Wilder JA (2011) Distribution and mechanism of α-amanitin tolerance in mycophagous Drosophila (Diptera: Drosophilidae). Environ Entomol 40(6):1604–1612. doi:10.1603/EN11136
Sturtevant AH (1921) The North American species of Drosophila. Carnegie Institution of Washington, Washington, DC
Sturtevant AH (1942) The classification of the genus Drosophila, with the description of nine new species. University of Texas Publication 4213:5–51
The Heliconius Genome Consortium (2012) Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 487(7405):94–98. doi:10.1038/nature11041
Throckmorton LH (1962) The problem of phylogeny in the genus Drosophila. University of Texas Publication 6205:207–343
Throckmorton LH (1975) The phylogeny, ecology, and geography of Drosophila. In: King RC (ed) Handbook of genetics, vol 3. Plenum Press, New York, pp 421–469
Toda MJ (2017) DrosWLD-Species: taxonomic information database for world species of Drosophilidae. Available at: http://bioinfo.lowtem.hokudai.ac.jp/db/. Accessed 25 Jan 2017
True JR, Edwards KA, Yamamoto D, Carroll SB (1999) Drosophila wing melanin patterns form by vein-dependent elaboration of enzymatic prepatterns. Curr Biol 9(23):1382–1391. doi:10.1016/S0960-9822(00)80083-4
Walker F (1849) List of specimens of dipterous insects of the collection of the British Museum. Part 4. British Museum (N.H.), London, pp 689–1172
Werner T, Jaenike J (2017) Drosophilids of the Midwest and Northeast. River Campus Libraries, University of Rochester, Rochester
Werner T, Koshikawa S, Williams TM, Carroll SB (2010) Generation of a novel wing colour pattern by the Wingless morphogen. Nature 464(7292):1143–1148. doi:10.1038/nature08896
West-Eberhard MJ (2003) Developmental plasticity and evolution. Oxford University Press, Oxford
Wittkopp PJ, True JR, Carroll SB (2002) Reciprocal functions of the Drosophila yellow and ebony proteins in the development and evolution of pigment patterns. Development 129(8):1849–1858
Yamaguchi J, Banno Y, Mita K, Yamamoto K, Ando T, Fujiwara H (2013) Periodic Wnt1 expression in response to ecdysteroid generates twin-spot markings on caterpillars. Nat Commun 4:1857. doi:10.1038/ncomms2778
Yassin A (2013) Phylogenetic classification of the Drosophilidae Rondani (Diptera): the role of morphology in the postgenomic era. Syst Entomol 38(2):349–364. doi:10.1111/j.1365-3113.2012.00665.x
Yeh SD, Liou SR, True JR (2006) Genetics of divergence in male wing pigmentation and courtship behavior between Drosophila elegans and D. gunungcola. Heredity 96(5):383–395. doi:10.1038/sj.hdy.6800814
Acknowledgments
We thank Toshiro Sekimura, Frederik H. Nijhout, and persons involved in the meeting at Chubu University in 2016 for stimulating us to write this chapter. We also thank Arnaud Martin and Takao K. Suzuki for reviewing this chapter, Masanori J. Toda for advice on taxonomy, Elizabeth Nakajima for English editing, and Noriko Funayama for hosting our research. A part of the writing was supported by KAKENHI (15K18586) and the Sumitomo Foundation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
This chapter is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
Copyright information
© 2017 The Author(s)
About this chapter
Cite this chapter
Koshikawa, S., Fukutomi, Y., Matsumoto, K. (2017). Drosophila guttifera as a Model System for Unraveling Color Pattern Formation. In: Sekimura, T., Nijhout, H. (eds) Diversity and Evolution of Butterfly Wing Patterns. Springer, Singapore. https://doi.org/10.1007/978-981-10-4956-9_16
Download citation
DOI: https://doi.org/10.1007/978-981-10-4956-9_16
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-4955-2
Online ISBN: 978-981-10-4956-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)